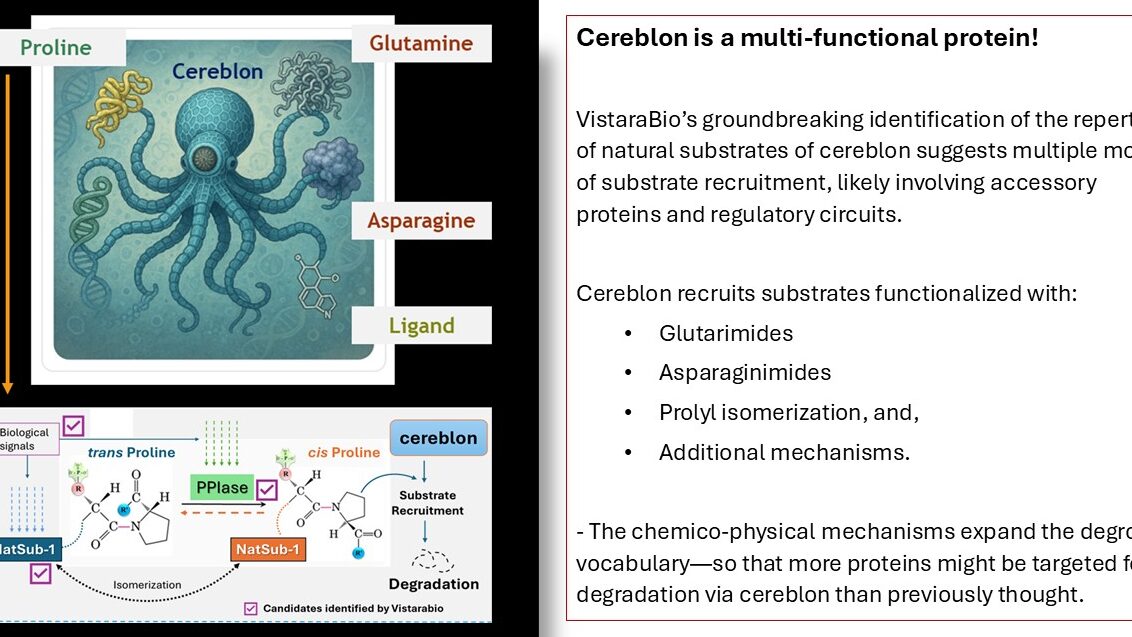
Multimodal substrate recruitment is a hallmark of cereblon function!
VistaraBio recently identified the repertoire of cereblon’s natural substrates [NatSubs] using its PINTAC process. Ongoing analysis has shown that the NatSubs can be bucketed into multiple subclasses of proteins based on their structures and amino acid signatures. These findings strongly suggest distinct, mutually independent substrate engagement modes by cereblon. The findings will spur highly insightful investigations leading to a better understanding of the roles of cereblon, including in thalidomide-based functions, resistance to the drug, and teratogenicity.
VistaraBio has designed targeted assays for each subclass of NatSubs which will report on the proteins and pathways involved with each subclass or candidate NatSub. Additional in vitro and cell-based assays developed at VistaraBio help screen for new chemical entities which provide high specificity, selectivity and reduced toxicity in preclinical models, as well as phenotypic effects in cell cultures or animals. Our assays combined with mass spectrometry observe compound-associated changes globally and map them into pathways and ascertain their mechanism of action.
Historically cereblon, a substrate recruitment factor for CRL4 E3 ligase, rose to prominence with the realization that it recruits and mediates degradation of certain proteins termed NeoSubstrates in presence of thalidomide or its analogs. Since then the protein has been a workhorse in the field of Targeted Protein Degradation, for applications involving molecular glue-like compounds, of which thalidomide is an example. NeoSubstrates are non-natural substrates of cereblon, which are distinct from the limited number of cereblon’s natural substrates known in the literature, and vastly expanded upon with VistaraBio’s approach.
Recently, cereblon has been shown by others to recruit natural substrates containing asparagine or glutamine amino acids at their C-termini [cN or cQ, respectively], which are included among the substrates identified by VistaraBio. Glutarimide analogs – compounds which bind to cereblon’s cQ substrates have been described for therapeutic applications in the future. VistaraBio has added a new aspect to cereblon substrate recruitment – prolyl isomerization of a subset of NatSubs, which likely involve the participation of peptidyl-prolyl cis-trans isomerases, [PPIases]. These expanded information and tools developed by VistaraBio, open up new areas in cereblon biology and the prospects of targeting cereblon with modalities which either trigger recruitment of select substrate categories, involvement of protein interactors, or regulate cereblon activity.
PINTAC expression provides a unique cell modification system!
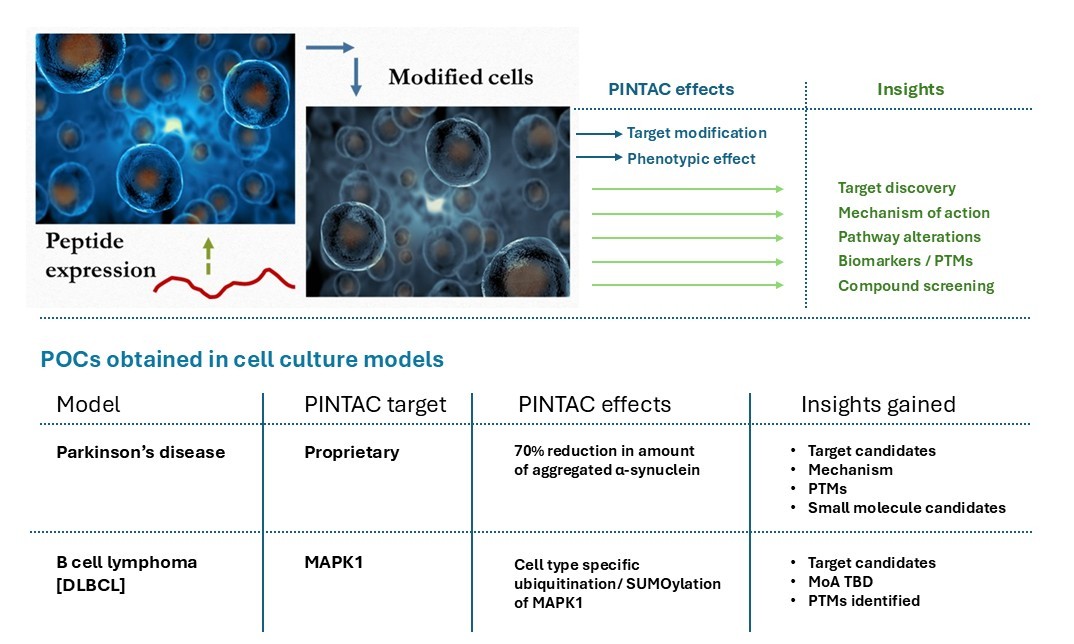
PINTACs are bioactive peptides by definition. They achieve highly selective modification of the intended target when expressed in cells which can be determined in protein preparations from the samples. The type of modification can vary with different targets based on several factors – the nature of the PINTAC-target interaction, their innate proximity in the cell, the cellular compartment in which the interaction occurs, characteristics of the cell type employed in the assay, and so on. On account of its robustness, some PINTAC peptides may give rise to a ‘null’ phenotype with respect to its target, likely by a squelching mechanism. The resulting cells can be employed in drug discovery research as transient ‘knock-outs’ which can further be tuned up or down with molecular biology. In either case, a target-dependent phenotypic effect is consistently observed which is determined with mass spectrometry.
VistaraBio has characterized over two dozen PINTAC peptides and determined proteomic alterations associated with them. The bioactivity of PINTACs has been determined with several PINTACs – a fist set in a toxin model of Parkinson’s disease [PD] and another in a B cell lymphoma model. The PD phenotypic screen identified compelling target candidates – the corresponding peptide candidates are being explored for development as therapeutic compositions. The associated information from this study also distinctly pointed out a new modality for PD therapeutics – restoration of function of damaged proteins.
PTMs will revolutionize biological discovery and the availability of novel therapeutic modalities!
VistaraBio has applied its synthetic biology approach for determining over 5,000 PTMs in nearly 1,500 proteins, many of which are of clinical interest. These proteins will be overproduced by VistaraBio for sales into research markets. Additional PTMs of proteins of client’s interest can be developed with VistaraBio’s PTM platform. VistaraBio is uniquely positioned for creating PTM maps in oncology or neurodegenerative diseases and has created assays for profiling them in singles or cohorts directly from disease samples. The tools include screening compound libraries against well characterized PTM-targets validated by VistaraBio. This platform will expand the biological and chemical spaces for new drug discovery, as well as generate new insights in large scale biology.
